IN CHAPTER 10, WE SAW that gene and genotype frequencies can vary considerably between different ethnic groups. In this chapter we will explore the underlying reasons for those differences. We will also see that our human activities can effect changes in the gene frequencies of other organisms, from bacteria to insects and other animals. More importantly, many of those changes in turn affect us.
The phrase “survival of the fittest” may conjure up an image of a big strong lion with his pride of lionesses producing many lion cubs. In that example, the genes of the strong lion, as well as those of the lionesses he chooses, will have a greater chance of being passed down to future generations of lions than the genes of a weak lion without any lionesses. We say that the genes of the strong lion are “selected” because of his ability to father more lion cubs. This, however, is not the only type of selection that exists in nature. Indeed, in the example of a field of four -o’clock plants in chapter 10, we saw a different type of selection. In that case, the preference of insect pollinators could increase the chances of one color gene being favored over another. Thus, selection is a process in which the individuals considered the “fittest” are those that have a higher chance of passing on their genes to the next generation. There are many ways to achieve this. To put it differently, the term “fitness” in this genetic sense does not necessarily refer to strong muscles or the ability to jog several miles. The fittest individuals are not necessarily the strongest, the smartest, or the healthiest. For example, it could be said that white four-o’clock flowers are less fit, not because they are sickly or smaller or have fewer flowers, but because they are not chosen by insect pollinators that prefer red flowers. In such a case, if insect pollinators prefer red flowers, they will not pollinate white or pink flowers, with the result that the white version of the gene will be underrepresented in subsequent generations. Therefore, the genes from the fittest individuals are those that are selected for in such a way that their proportion will increase in successive generations. The genes from less fit individuals are selected against and their proportion will decrease in successive generations.
Selection can either be natural or artificial. Natural selection takes place in a natural environment, and selective forces there can be any number of factors such as drought, high or low temperatures, existence of predators, insect pollinators, and so on. Individuals genetically well-adapted to these conditions, or well-adapted to changing conditions, can survive and proliferate. Genetically less-adapted individuals will have a harder time and may well become extinct. How does selection by natural or artificial means take place? Possibilities for selection of some individuals exist because not all individuals in a population are genetically identical and equally fit. This is because each individual, as we saw, is the product of gene shuffling during meiosis, and therefore different individuals do not possess the same gene combinations.
Selective (artificial) breeding for desired characteristics, along with natural selection imposed by the environment, have produced our domesticated animals, pets, and crops. Successful selective breeding also requires variations within populations. That is, if all the animals or plants had the same characteristics, ones that produced more milk or laid more eggs or had higher oil content—or whatever qualities we wanted to enhance—would not exist. Thus there can be no selection without variation in populations of animals and plants. The ultimate source of this variation is mutations. Let us now consider a few examples of artificial selection.
Humans have produced many successful strains of domesticated animals and crops. These animals and plant have the highest quality of the desirable properties humans look for. For example, if one observes the milk goats at a local county fair, one sees that they all have tremendously large udders, so large that they would have difficulty walking if let out into the mountainous terrain of their ancestors. These goats were bred for the purpose of high milk production and originated from goats that produced more milk than average. Over time, breeders selected progenies from these goats with even higher milk production and bred them again, until the milk production was much higher than in the original goats, and, by consequence, large udder size was obtained. This example shows that artificial selection by breeders can go against what would have been selected for in a natural environment. In nature, goats with inordinately big udders would surely be selected against, as they would not have the agility of small-udder goats and would be at a disadvantage. Genes responsible for large udders would then quickly be diluted in a population of wild goats.
Thanks to the “Green revolution,” the yield of our grain crops, such as rice and wheat, has increased tremendously. Here also, selective breeding of individuals showing desirable characteristics was involved. In some cases, mutants obtained through the use of mutagens were used as progenitors of the high-yield varieties in current production. Artificial selection is generally beneficial to human consumers, but it can sometimes have unforeseen, deleterious consequences.
For example, in our efforts to produce animals and plants with just the desired characteristics, our domesticated animals and crops now have limited genetic diversity. This is due to the fact that, by selectively breeding a limited number of individuals, we stop propagating those individuals that do not possess the characteristics we desire. The rejected individuals and the genes they carry are then at risk for extinction. However, the individuals we select against may possess characteristics of which we are unaware. For example, if new diseases or pests of crops arise, it may well be that the rejected individuals naturally possess genes making them resistant to these pests and diseases, while the selected individuals do not. In other words, through selective breeding, we may have eliminated from the gene pool traits that may be important at some point in time under different conditions. This lack of genetic diversity can make the entire crop susceptible, and the disease or pest may quickly devastate it.
This is what happened with the Irish potato famine of the 1840s, in which fungi attacked the potatoes. All the potato strains in Ireland were similar and did not have any natural defense again the fungi, and the potato crop was wiped out for several years. In the 1970s in the United States, the corn crop was similarly destroyed because almost all the U.S. corn was susceptible to southern corn leaf blight, another fungal disease.
Similar problems associated with limited genetic diversity can arise in the breeding of pets. Professional breeders are careful to avoid breeding closely related dogs in order to maintain genetic diversity. Nevertheless, for popular breeds of dogs in which there are pressures to produce many puppies, medical problems such as hip dysplasia and behavioral problems have crept into certain breeds. Thus, animals and plants with highly desirable characteristics and limited diversity of other genes may encounter unexpected difficulties. Let us now study a case of natural selection in human populations.
Selection can occur on Mendelian, or single-gene, traits, but also on traits determined by many genes, since selection acts on the phenotype, as we have seen in the case of goats, lions, and four-o’clock plants. An example of a trait determined by many genes is skin color in humans. Natural selection for or against skin pigmentation is well understood and took place tens of thousands of years ago as modern humans left Africa to colonize the rest of the planet. People from tropical areas with high exposure to the sun are darker in order to protect themselves from the dangers of ultraviolet rays. But ultraviolet rays also serve a necessary function by helping chemicals present in the skin to convert to vitamin D. People with dark skin still make enough vitamin D because high sunlight intensity in the areas in which they live compensates for lower efficiency due to skin pigmentation. On the contrary, individuals from higher latitudes and areas with greater cloud cover have extremely light skin. These individuals use what little ultraviolet light they do get for the purpose of producing vitamin D. You now understand why human variants with light skin pigmentation were not fit to survive and proliferate in the tropics: the harmful effects of ultraviolet light selected against them. In a similar fashion, humans with dark skin pigmentation were at a disadvantage in higher latitudes because they could not make enough vitamin D to stay healthy.
Today, unlike in ancient times, people travel easily throughout the world and carry in their genome the genetic history of their ancestors. Our genes do not change just because we are now in a different geographic area or environment. Light-skinned individuals can now survive in tropical climes because they wear protective clothing and use sun screen. Dark-skinned individuals can now survive in northern climes because they obtain vitamin D from food. Thus, our genetic heritage is a reflection of all the selective pressures that our ancestors have encountered.
Let us imagine a mutation in an insect that made it lose its wings. How would this affect its fitness? We might expect that this would be bad, that is, an insect without wings would have lower fitness compared to the same insect with wings. But what if loss of wings happened to insects that were blown off shore, onto an island, in an area that is often windy? The island is small and the area windy, so it might be safer for insects to walk around rather than try to fly and get blown away into the ocean. Thus whether a particular phenotype is higher or lower in fitness depends upon the environment or circumstances in which an individual lives.
The case of insect pests demonstrates the way in which human activity changes the environment to affect selection. Various weevils thrive on agricultural crops and destroy them. Populations of weevils have all types of natural genetic variations because they breed freely in the wild and accumulate random, spontaneous mutations. But farmers started using pesticides to get rid of these noxious pests. When a field is sprayed with pesticides, weevils with natural resistance have higher fitness than weevils without natural resistance to the pesticide. Thus most of the susceptible weevils are killed, and a few weevils, those with natural resistance to the pesticide, survive. What happens after the effects of the pesticides have worn off? Susceptible weevils from surrounding areas that were not sprayed enter the field since there is plenty of food now and fewer weevils to compete with for that food. The net effect of the susceptible weevils “migrating” in from the surrounding fields is to swamp the gene pool with genes susceptible to pesticides. Thus, resistant weevils are again in a minority.
But now consider a field of genetically modified crops that make Bt toxin all the time in the plant tissues themselves (see chapter 6). Unlike the situation above, there is no way for the weevils to get away from the pesticide because it is always present in the crops. Even if more weevils migrate in from the surrounding areas, those susceptible are killed continuously, and, as a result, only resistant weevils survive. These survivors mate and have more weevils. Soon, a higher percentage of weevils have pesticide-resistance genes. There are insufficient numbers of weevils with susceptibility genes surviving to dilute those with resistance genes. Among this large number of resistant insects, some, by random chance, develop even higher resistance to Bt toxin. Of course, those with higher resistance will continue to increase in number over those with less resistance as long as the Bt toxin is present. In the end, only resistant weevils are left, and the pesticide has lost its effectiveness.
This is the scenario that quickly started to unfold with genetically modified Bt crops. Indeed, very soon after the commercial release of Bt-modified crops, farmers began to notice that there were a lot of insect pests on their Bt crops that were not supposed to be there. These insects had become resistant to the Bt toxin. In order to prevent a massive proliferation of Bt-toxin-resistant insect pests, farmers that plant Bt-modified crops must now agree to plant a percentage of their acreage with non-Bt-modified crops close to the field of Bt-modified crops. The acres of non-Bt-modified crops provide refuge for insect pests that are susceptible to Bt-toxin. Without this refuge, the genes for Bt-toxin susceptibility will be eliminated from the population, and Bt toxin, whether sprayed on the fields or made by the crops themselves, will no longer be effective.
All living things are faced with selective forces. In the case of bacteria, human use of antibiotics is a major selective force. We presently have a whole arsenal of antibiotics to fight bacterial infections. But until the 1940s there were no antibiotics at all! Antiseptics such as alcohol and iodine were used well before antibiotics were discovered, but they were not effective against diseases like tuberculosis, pneumonia, and other internal or deep bacterial infections. In the pre-antibiotic days many women chose not to deliver their babies in hospitals because many babies and women died of bacterial infection. Hospitals were particularly unsanitary because doctors easily transmitted bacterial infections among their patients, in spite of all the precautions taken. In World War I, for example, more casualties were due to infections in wounds than to the wounds themselves because there were no antibiotics.
The discovery and subsequent production of penicillin in the 1930s and 1940s dramatically changed the prospect for patients with bacterial infections. Antibiotics became the magic bullet by which bacterial infections could be stopped in their tracks. The 1945 Nobel Prize for Medicine was awarded to Sir Alexander Flemming, Sir Howard Walter Florey, and Ernst Boris Chain for their life-saving work, the discovery and industrial production of the very first antibiotic: penicillin.
So, what has this got to do with selection? Antibiotics provide a tremendous selective force against bacteria, because the function of antibiotics is to kill bacteria. Only those bacteria that can somehow survive an onslaught of antibiotics pass on their genes to their progeny. This can only happen if some bacteria become resistant to an antibiotic. How is this possible? Bacteria have a variety of ways to thwart the action of antibiotics. First, an antibiotic cannot work if it cannot remain inside the bacterial cell. Some bacteria can actively pump out antibiotics and thus survive. Other bacteria can chemically break down the antibiotic so that it is no longer active. Finally, the target of the antibiotic, which is often a bacterial protein, can be modified (mutated) so that the antibiotic can no longer recognize it and poison it. With all these different ways—that is, mutations in different genes—to get around the killing effects of antibiotics, it is no surprise that resistant bacteria arose soon after their introduction.
Penicillin, for example, affects the protective cell wall of a group of bacteria. When cell-wall formation is blocked by the action of penicillin the bacteria burst and die. In the 1940s when penicillin was first introduced, 100 percent of staph infections were effectively treated with penicillin. Now, greater than 90 percent of staph bacteria are resistant to penicillin. A newer antibiotic called methicillin was introduced to counter the rise of penicillin resistance. In 1974, only 2 percent of staph infections were methicillin resistant, but, by 1997, 50 percent of the staph infections had become methicillin resistant. In 2000, over 30 percent of the most common bacteria that cause ear infections in children had developed resistance to penicillin. A similar trend has been found with almost every antibiotic produced to date. So there is an escalating arms race between the bacteria and us. We must continue to develop new antibiotics to keep ahead of the resistance that bacteria inevitably develop.
Why has so much antibiotic resistance appeared? To answer this question, we need to think about selection and survival of the fittest. As patients taking antibiotics, we are exerting selective pressure for antibiotic resistance among the bacteria that infect us. If we take a properly high dose of antibiotics for a sufficient amount of time, chances are excellent that the bacteria causing the problem are completely eliminated. Problems begin when we take an insufficient dose or an incomplete series. Under these circumstances, a few bacteria that survive better than the average, thanks to mutations, are not killed because of the lower dose or shorter time involved. They can now grow up and provide a large number of bacteria that can develop even higher resistance through mutation. These bacteria survive in our bodies and can also find their way into our environment through coughing, sneezing, and defecating. Now if we try to fight this bacterial infection with the same antibiotic, it is no longer as effective as before. The resistant bacteria also can infect other human victims. Then, if these individuals use the same antibiotic, all of the invading bacteria will be resistant to that antibiotic. Consequently, the antibiotic will no longer be effective for the population.
Overuse or inappropriate use of antibiotics also contributes to the rise of antibiotic resistance in bacteria. For example, using antibiotics for viral infections does nothing for the infection because viruses are not susceptible to antibiotics. Yet this course of action can promote antibiotic resistance of harmless bacteria that normally reside in our bodies. In and of itself, this does not seem particularly dangerous. However, these antibiotic-resistant harmless bacteria can transmit their genes to pathogenic bacteria as we see below, thereby rendering them resistant as well.
Finally, tremendous amounts of antibiotics are used in agriculture. Antibiotics are used to treat bacterial infections and promote growth in the beef and poultry industries. Antibiotics are also used for plant bacterial diseases. Thus, not surprisingly, bacteria resistant to widely used antibiotics have been found on farms. The particular problem with harmless bacteria or bacteria that are plant pathogens is the way that some bacteria have developed antibiotic resistance. Recall that many bacteria contain a minichromosome, or plasmid (chapter 5). These plasmids can be spontaneously transmitted to different bacterial species. Scientists have discovered a particular type of plasmid that they have named “multidrug-resistant plasmid.” This plasmid has several different genes that confer resistance to a number of antibiotics. It can be transmitted from a harmless bacterium to a disease-causing one or from a bacterium that only infects fruit crops to one that is harmful to animals and people. This means its resistance to multiple antibiotics can be transferred to disease-causing bacteria. By all these different means, the rise of antibiotic resistance has reached epidemic proportions. If this situation is not dealt with effectively, we may soon be back to the pre-1940s days when we were without an effective weapon against bacterial infections.
We saw in chapter 10 that selection has little effect on recessive disease traits because carriers do not exhibit the trait and therefore are not selected against. For example, in the case of PKU, 99 percent of the PKU genes are found in heterozygote carriers who are phenotypically normal. This means that it takes an extremely long time for recessive traits, even lethal ones, to be eliminated from the population. Indeed, as the recessive form of the gene is eliminated in the form of unfit, homozygous affected individuals, a greater percentage of the disease gene is carried by phenotypically normal heterozygous individuals. Was there a reason for such disease traits to appear in fairly high numbers to begin with? Why are there differences between different ethnic groups for these and other diseases? We are finding out that the answers to these two questions reside in the fact that for many recessive diseases there is a heterozygous advantage to carrier individuals.
The classic case is that of a disease we already mentioned in chapter 3, sickle-cell anemia. Recall in the last chapter that the incidence of sickle-cell anemia is much higher (about 1 in 400) among African Americans than among Caucasian Americans, who show a frequency of around 1 in 2,500. Why is there such a disparity between these two ethnic groups? The answer lies in the genetic history of these two populations. African Americans came from tropical regions with high incidence of malaria, whereas most Caucasian Americans came from temperate regions in Europe, where malaria was not prevalent.
Individuals who are heterozygous for the sickle-cell trait do not, of course, exhibit sickle-cell anemia because the trait is recessive. However, these heterozygous individuals, it turns out, are more resistant to malaria. Malaria is a microbial infection spread by mosquitoes that carry the malaria parasite. When mosquitoes carrying parasites bite a person, the parasites enter the blood stream and the red blood cells of this individual. The parasites infecting the red blood cells make the inside of the cells acidic. This does not affect normal β-hemoglobin in the red blood cells of a person homozygous for the normal β-hemoglobin gene. As a result, the red blood cells of this person retain their normal shape, and infection by the parasite can continue. Death results if the infection is left untreated. However, if the red blood cells of an individual heterozygous for the sickle-cell trait becomes acidic, the red blood cell takes on a sickle shape due to the mutated β-hemoglobin. The abnormal shape of the red blood cells signals the body to eliminate them. Thus, the malaria infection cannot proceed and the infected person survives.
This mutation, then, in the heterozygous state, provides a selective advantage to people living in areas where malaria is prevalent. Of course, it does not provide any advantage to people who do not experience malaria. The sickle-cell trait is debilitating to homozygous individuals and is thus a selective disadvantage. The selective advantage it provides to heterozygous individuals offsets this disadvantage and maintains a higher proportion of sickle-cell trait in the population than one would otherwise expect. This is a classic example of heterozygous advantage that maintains a disease gene in a population.
More recently, a mutation in another gene called G6PD (glucose-6-phosphate dehydrogenase), which codes for an enzyme necessary for red blood cells to obtain energy from glucose, has also been found to exhibit heterozygous advantage. Certain mutations in G6PD, which in the homozygous state causes anemia, also provide resistance to malaria infection in the heterozygous state. Similarly, mutations in other genes for proteins found in red blood cells are thought to confer selective advantage in resistance to malaria. Malaria may have been quite prevalent for a long time and thus a strong selective force acting on the genetic makeup of peoples from tropical regions.
A last example of heterozygous advantage is that of PKU, a disease you have heard about already in chapters 3 and 10. As with sickle-cell anemia, homozygous, recessive individuals are highly disadvantaged. However, again as with sickle-cell anemia, scientists postulate that the PKU trait provides a selective advantage to heterozygous individuals. The mild, damp climate of the British Isles is conducive to the growth of molds in grains and other stored foods. These areas also suffered repeated widespread famines. When one is starving, moldy food is better than no food. Molds have toxins that, among other things, cause spontaneous abortion. However, women heterozygous for the PKU trait appear to have had a lower spontaneous-abortion rate than women homozygous for the normal gene. Thus, the presence of the PKU trait in a heterozygous condition could have favored the survival of offspring from heterozygous women who resorted to eating toxic, moldy food. This would have favored the persistence of the PKU gene in populations where a combination of periodic starvation and moldy foods existed.
As we understand more about the evolution of genetic traits in humans, other recessive genetic diseases for which there is heterozygous advantage may be recognized. For example, cystic fibrosis is another disease that is postulated to confer heterozygous advantage in areas with diarrheal diseases.
We have seen that deleterious traits can remain in the population if they are recessive and if they also provide heterozygous advantage. But we also know of dominant genetic diseases. How do these conditions appear and remain in the population? Dominant traits initially appear as do all other genetic variations: by mutation. In fact, some dominant genetic diseases show a high rate of new mutations; that is, they are not inherited from the parents but appear anew. For example, Marfan syndrome is a dominant genetic disease caused by a mutation in a connective-tissue protein called fibrillin. The gene encoding this protein is about 200,000 base pairs long, quite a large gene. Because most of this protein’s structure is important for its proper functioning, the large gene that codes for it provides a large target for mutations. It is estimated that approximately 25 percent of Marfan cases are due to new mutations.
Scientists are identifying more dominant mutations as time goes by, but the great majority of these mutations are extremely rare. Some mutations are probably not recognized as dominant because the individual with the dominant mutation does not survive to pass it on. The first dominant mutation found in humans, brachydactyly, was recognized as such because of one large family affected with it. Similarly, other rare genetic diseases are only recognized when a large family with a particular phenotype is recognized and pedigree analysis is performed.
Yet we find that some dominant diseases are more common than one might expect from a rare mutation and a few families passing on this trait. This is the case for Marfan syndrome, Huntington’s chorea, and familial early-onset Alzheimer’s disease. The three diseases share a common feature: individuals with the disease traits do not express the disease phenotypes until they are past their reproductive age. That is, by the time the phenotype appears, individuals may have already passed on their defective genes to the next generation. Thus, selection does not have a chance to act because affected individuals reproduce before the disease is apparent. Selection can only act on the phenotype, and selection manifests itself in individuals’ success or failure in passing on their genes.
We saw already that, depending upon the genetic history of different peoples, the frequency of genetic traits can differ dramatically among different populations. This difference among groups is accentuated by small population size. Two phenomena, the founder effect and random genetic drift, account for the effects of small population size on gene frequency.
The founder effect is so named because it is observed when a small group of individuals founds a new population. This effect can also be observed when there is a sudden reduction in population size. To illustrate this concept, imagine a very rare genetic disease with a frequency of 1 in 100,000 people. Now, let us imagine that from this population, 100 people decide to emigrate to an unsettled part of a territory. Let us further imagine that, just by chance, the one individual with the disease gene is among the hundred individuals who leave the group. So, just by random chance, the frequency of disease in this new population jumps to 1 in a 100 instead of 1 in 100,000! Thus, without selection, since no one died before passing the gene on, all of a sudden just by subdividing into a small population, the rare trait is now not so rare. This is called the founder effect because it is particularly evident when a founding population colonizes a new land. However, the effect is no different if the population was suddenly subdivided or decimated. For example, assuming the same frequency of 1 in a 100,000, if a huge volcano erupted and almost everyone was killed except the same 100 as above, we would again end up with a 1 in 100 frequency.
DNA sequencing has been used extensively in the study of human origins and evolution. By looking at DNA sequences from different organisms and measuring how much these sequences differ, it is possible to determine the time of appearance of a species. The rationale is that variations in DNA sequences are due to mutations, and the longer the period of time that passes, the more mutations that can accumulate. By comparing the DNA sequences of homologous genes obtained from various individuals belonging to different groups, we can measure how different the DNA sequences are and thus how much time has elapsed since the two lineages split. That is, sequence comparisons allow scientists to estimate the time when the two groups shared a common ancestor.
One particularly convenient piece of DNA to sequence for determining evolutionary relatedness among humans is mitochondrial DNA. In addition to nuclear DNA, animal cells contain DNA in bodies called mitochondria. Mitochondria are the “energy factories” of cells and contain a bit of genetic material. Because mitochondrial DNA is relatively short—about 16,600 base pairs in humans—it is easier and faster to sequence than the whole nuclear genome. Also, portions of mitochondrial DNA mutate more rapidly than many of the genes in our nuclei, in a time span of a few thousand years.
Mitochondrial DNA from various human ethnic groups was sequenced, and these sequences were used to reconstruct an evolutionary tree. The tip of the branches represents the ethnic groups living today. The groups that branch out closer to the base of the tree (the root) represent the earliest divergences of the human ethnic groups. It turns out that people from sub-Saharan Africa appear at many different branches of the tree. A group consisting of only sub-Saharan Africans branch first from people from other continents. One interpretation of this result is that modern human beings, Homo sapiens sapiens, first appeared in Africa about 150,000 years ago (Figure B.11.1). These humans subsequently migrated out of Africa and colonized the whole world. There are more different DNA sequences from people in Africa than elsewhere on earth because humans were in Africa first. Interestingly, mitochondrial DNA is inherited only from mothers. An egg contains lots of mitochondria, while the sperm head that fertilizes the egg does not. This has led to the intriguing concept of “Eve,” the first group of African women whose mitochondria is still (with base pair changes) within all of us. “Eve,” however, should not be considered a single woman; rather, there probably were thousands of “Eves” that gave rise to modern humans. Nevertheless, mitochondrial DNA sequences fit well with the idea of a founder effect at the root of the human evolutionary tree.
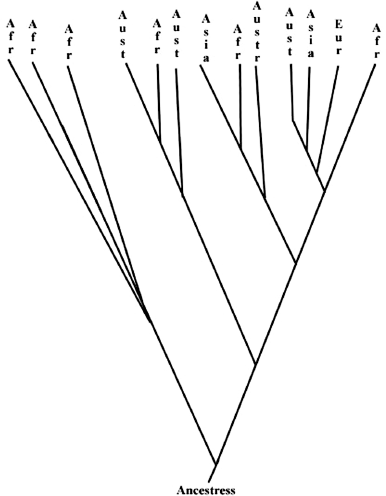
Figure B.11.1 Simplified Evolutionary Tree of Humans. This evolutionary tree of humans was derived from the study of mitochondrial DNA by Allan Wilson. The key features include a group of Africans branching out from near the root of the tree; Africans are also represented on all the other branches.
DNA sequence comparisons were also made on the DNA from the Y chromosome. As we know, this chromosome is found only in males and thus transmitted only through fathers. Studies of the differences in sequence on the Y chromosome among different ethnic groups are similar to those obtained with mitochondrial DNA. Here also, scientists have interpreted this evidence as African “Adams” from whom humankind is derived. These “Adams” and “Eves” migrated out of Africa to Europe and Asia and finally to the Americas. The thousands of years during which the first humans traveled to all corners of the world allowed for natural selection and genetic drift to occur. Natural selection worked at the level of skin color and general body build. Random genetic drift and founder effect explain DNA sequence variations that do not seem to have selective advantage or disadvantage. Thus, Y-chromosome studies also support an African founder effect for the human family.
In addition to revealing the origins of Homo sapiens, mitochondrial DNA sequencing may shed some light on the fate of an extinct branch of the human family. It is known that the modern humans that colonized Europe and the Middle East (they are often called Cro-Magnons, after the place in France where their fossils were first discovered) encountered other humans, the Neanderthals (Homo sapiens neanderthalensis). These Neanderthals were well adapted to the cold climate prevalent during the last Ice Age. However, as the climate warmed up, Cro-Magnons progressively took over the Neanderthals. The latter became extinct about 30,000 years ago. A question that has intrigued paleontologists is how Neanderthals became extinct. Two theories are proposed: one that Cro-Magnons eradicated Neanderthals through warfare, while the other theory holds that Neanderthals and Cro-Magnons interbred, to result in today’s European and Middle Eastern populations. Researchers were able to isolate mitochondrial DNA from a limited number of Neanderthal fossil bones. Sequencing this DNA indicated no close relationship between modern human mitochondrial DNA and Neanderthal mitochondrial DNA. This seems to rule out the interbreeding theory. Therefore, it may well be that our ancestors committed genocide on the Neanderthals to occupy the ecological niches previously held by them.
Random genetic drift is a process that occurs in a small population after the founding of that population. As the name indicates, the process is random, that is, it is not due to a fitness advantage or disadvantage. We will demonstrate this with a real-life example of the group represented by Old Order Amish Mennonite Church. This group arose in central Europe during the seventeenth century and began migrating to the United States in the early eighteenth century, predominantly to Pennsylvania and Ohio. Those who espouse the traditional teachings of this church wear plain clothing: men wear hats and do not trim beards, while women wear long dresses, capes, and bonnets. They also use horses and buggies rather than automobiles and generally shun modern conveniences. These people stay in relatively small communities and tend to socialize within their groups. The necessary ingredients for observing random genetic drift is present in this situation, a small founding population that tends to marry within its group. By emigrating in small numbers, the group has set up a founder effect. For example, the frequency of dwarfism, a recessive trait, in the general population is roughly 1 in a million. By using the Hardy-Weinberg law that we learned in chapter 10, we can calculate that in the general population there are roughly two in 1,000 who are carriers (figure 11.1.A). However, just by random chance, two carrier heterozygotes were present in a founding population of one hundred Amish emigrants. So, the founder effect increased the frequency of dwarfism carriers from two in a 1,000 to two in 100 in this particular emigrant group. Using the frequency of carriers of two in 100 we can calculate how many homozygous dwarf individuals we expect to find in this Amish village (figure 11.1.B). This frequency is expected to produce 1 dwarf in 10,000 people. Note that this number is much higher than 1 in 1,000,000, as is found in the general population. However, we actually know that in this particular Old Order Amish community the frequency of dwarfism is 1 in 14. This indicates that the frequency of dwarfism gene in this community is higher today than it was when the population was founded. From this, can you calculate the expected frequency of carriers in the population?

Figure 11.1 Frequency of Genes and Genotypes in a Small Population. A. In the general population, dwarfism is quite rare and found in 1 out of a one million, or 10-6, the frequency of aa, since dwarfism is a recessive trait. We can calculate the frequency of a, the dwarf form of the gene, as 10-3 and the frequency of carriers as 2 in 1,000. B. We calculate the dwarfism gene frequency in an emigrant population of 100 that just by chance has two heterozygotes (Aa) among them by putting 1 in a 100 or 0.01 in both the boxes for the heterozygous carriers. We see that 0.01, or 1 percent, is the frequency of the a gene in this smaller population. C. We now can calculate the expected frequency of dwarfs (aa) in this new population as 0.0001 or 10-4. That is, we expect that with the dwarfism gene’s frequency of 1 percent, there will be 1 dwarf in 10,000 births. D. Now we use the actual frequency of dwarfism observed in this population, 1 in 14, which is 0.07. By finding the square root of 0.07, we get 0.27, or 27 percent, as the estimated frequency of the a gene in the population. Note that this is significantly higher than the 1 percent frequency of the a gene observed in the founding population. This is genetic drift: that is, just by random chance in a small population, the frequency of a deleterious gene can actually increase. Note that this Punnett square is incomplete because the frequencies of A, AA, and Aa are not calculated.
We can use the Hardy-Weinberg law to calculate the frequency of the dwarfism gene in this community, which turns out to be approximately 25 percent (figure 11.1.C). How did this happen? Clearly, dwarfism, especially for a group of people who do not believe in using modern conveniences, is not advantageous. This increase in dwarfism gene was due to random genetic drift. Just because the population size was small, random effects changed the frequency of genes, even genes that might have a selective disadvantage. Indeed, an extensive study of a number of Amish communities shows an unusually high incidence of different genetic traits, not just dwarfism. That is, an unusually high frequency of genetic traits that are rare in the general population exists in the Amish community, for example, traits such as the neuromuscular disorder called Amish nemaline myopathy, and Amish brittle hair syndrome.
Founder effect and random genetic drift affect all living creatures that experience a dramatic decrease in population size. For example, it is postulated that the massive hunting pressure that reduced the population size of lions may have contributed to the loss of fertility among these animals. It is expected that other endangered species reduced in number by loss of habitat or hunting will face these same random effects of small population size. Thus, even deleterious genes may increase in frequency in populations that are already in danger, just based on the random nature of passing on genes in a small population.
In the context of genetics, fitness means the ability to pass on genes to the next generation. This may be due to the strength and fecundity of some individuals, but it can also be the result of attraction to mate or pollinator. Selection is the force that favors or disfavors some phenotypes relative to others under a certain set of environmental circumstances. For selection to act, there must be variation of traits within a population. Without this genetic diversity a population is ill prepared for changes in its environment. Selection can be natural or artificial. Artificial selective forces such as antibiotics and pesticides can quickly increase resistance among bacteria and insects. Human genes reflect the history of the selective forces experienced by our ancestors. For example, some deleterious recessive genetic traits exist in higher proportion than expected due to the heterozygous advantage they confer. Also, some diseases are more prominent than expected, not because of any selective advantage, but because of the founder effect or random genetic drift. Thus, small populations, just because of their size, can contain a high proportion of less-fit individuals.
1. Founder Effect
The founder effect can be demonstrated in a fun, tasty way. Any individual-size packages of candy pieces, such as small bags of M&Ms, will work. The bags of candy must have different colors of candy pieces and have relatively few pieces in each individual bag.
Now, open one package and count how many pieces there are of each color of candy. You may eat all the candy once the numbers have been noted. Then do this with a few more bags of the candy. Avoid the danger of this exercise by having your friends and family members help out, at least in eating the candies! At this point, tally the number of different colors from each bag. Now estimate the ratios of the different colors of M&Ms at the factory by adding the number of each color of M&Ms in all the bags together and determining their ratio. Now compare this ratio for the "factory" with the ratio of different colored M&Ms in each individual bag.
This exercise demonstrates what is known as the founder effect in population genetics. Imagine that the M&Ms represent individuals in a population with different colors representing different forms of a gene. In the large population, at the M&M factory, each color is represented by many individuals. Yet when they put just a few M&M into each of these bags, just by chance more of some colors and fewer of others are put in the bag. Each bag has a different random number of colors. Similarly, just by chance in a population, a rare genetic disease trait may appear.
2. Random Genetic Drift
This next exercise demonstrates random genetic drift. As with the founder effect, drift is more evident with small populations because random chance has greater impact on small populations than on larger ones.
Begin with a large number of pieces of colored paper, let’s say green, purple, and orange. We suggest at least 100 pieces, with 60 percent green, 20 percent purple, 20 percent orange. This is your original "genetic pool," wherein each different form of the gene is represented by a given color and each piece of paper represents an individual.
Then pick ten pieces of paper randomly (that is, blind, without regard to color) from this initial "genetic pool." This is generation 0. Keep track of how many of each color you have at each generation. Note that the chances are that you did not get the same proportion of colors as in the original "genetic pool" of six green, two purple, and two orange. This, again, is the founder effect. That is, just by chance, you might pick many more purple or orange pieces than is representative of the original pool, but, just as likely, you might not get any purple or orange.
Now, we will reproduce this population for a few generations. At "each generation," randomly pick a piece of paper from your small population of ten. After you pick, note the color, then return the piece of paper to the population of ten, mix the pieces of paper, and then pick another and note its color. Do this ten times so that you again have ten individuals. This process simulates the genes that are randomly passed onto the next generation. The number of different colors represented by these ten individuals may or may not be the same as the original 10 pieces. But now, these ten colors you picked will form generation 1. If the proportion of colors is the same as generation 0, just use the same pool of ten individuals to make a new generation. If the proportion is different, then get more of the colors you need to form a new generation representing the colors you picked. This is generation 1, from which you will again pick ten individuals to form generation 2 and so forth until you have ten generations.
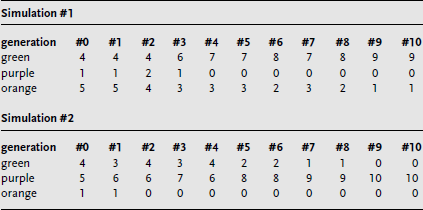
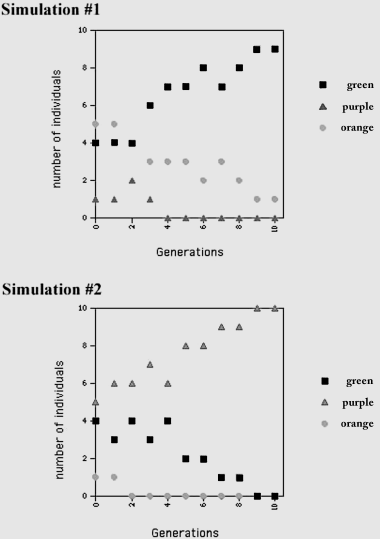
What you have done is simulate ten generations for a very small population of ten individuals. To visualize the result, you can plot the number of pieces of the three different colors on the y-axis of a graph as a function of the generation number on the x-axis, as shown for the two sample simulations in figure B.11.2 and in table 11.1. What happened to the numbers of different colors in your simulation? If this simulation was repeated, what might you expect?
Note in the example shown that sometimes one color can become predominant or even become the only color in the population. That is, if one color is eliminated, there is no way for that color to reappear in the population without mutation (which occurs at a very low rate) or migration. If this happened in a real population, it would mean that genetic diversity would dramatically decrease. A gene trait that is not highly represented in the original population can be eliminated, as in simulation #1 with purple, and even traits that are prevalent in the original population can disappear, as with green in simulation #2. Conversely, a rare trait can be the only one in the population, as with purple in simulation #2.
Table 11.1 Results from Two Simulations of Random Genetic Drift
Figure B.11.2 Simulations of Random Genetic Drift. Each form of the gene is represented by colors of paper represented by different symbols on the graph. The frequency of different forms of the gene at each generation is shown. Though we began with the same proportion of the colors, just by random chance, the proportions of purple and orange are different. This is the founder effect. Then with each generation, because the population size is small, by random chance certain colors can predominate in the population or even be the only color in the population.